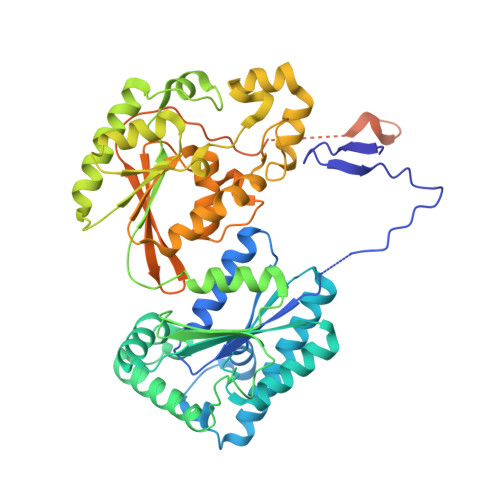
A Direct Substrate-Substrate Interaction Found in the Kinase Domain of the Bifunctional Enzyme, 6-Phosphofructo-2-kinase/Fructose-2,6-bisphosphatase
Kim, S.G., Cavalier, M., El-Maghrabi, M.R., Lee, Y.H.(2007) J Mol Biol 370: 14-26
- PubMed: 17499765
- DOI: https://doi.org/10.1016/j.jmb.2007.03.038
- Primary Citation of Related Structures:
2DWO, 2DWP, 2I1V - PubMed Abstract:
To understand the molecular basis of a phosphoryl transfer reaction catalyzed by the 6-phosphofructo-2-kinase domain of the hypoxia-inducible bifunctional enzyme 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase (PFKFB3), the crystal structures of PFKFB3AMPPCPfructose-6-phosphate and PFKFB3ADPphosphoenolpyruvate complexes were determined to 2.7 A and 2.25 A resolution, respectively. Kinetic studies on the wild-type and site-directed mutant proteins were carried out to confirm the structural observations. The experimentally varied liganding states in the active pocket cause no significant conformational changes. In the pseudo-substrate complex, a strong direct interaction between AMPPCP and fructose-6-phosphate (Fru-6-P) is found. By virtue of this direct substrate-substrate interaction, Fru-6-P is aligned with AMPPCP in an orientation and proximity most suitable for a direct transfer of the gamma-phosphate moiety to 2-OH of Fru-6-P. The three key atoms involved in the phosphoryl transfer, the beta,gamma-phosphate bridge oxygen atom, the gamma-phosphorus atom, and the 2-OH group are positioned in a single line, suggesting a direct phosphoryl transfer without formation of a phosphoenzyme intermediate. In addition, the distance between 2-OH and gamma-phosphorus allows the gamma-phosphate oxygen atoms to serve as a general base catalyst to induce an "associative" phosphoryl transfer mechanism. The site-directed mutant study and inhibition kinetics suggest that this reaction will be catalyzed most efficiently by the protein when the substrates bind to the active pocket in an ordered manner in which ATP binds first.
Organizational Affiliation:
Department of Biological Sciences, Louisiana State University, Baton Rouge, LA 70803, USA.